Welcome to Atomap’s documentation!
News
2025-1-18: Atomap has moved to GitHub
The development of Atomap has moved from https://gitlab.com/atomap/atomap to https://github.com/atomap-dev/atomap. While GitLab has been a good home for Atomap, most of the HyperSpy community is on GitHub. So this move will make collaboration with the rest of the community easier.
2025-1-5: Atomap 0.4.2 released
This is a small release which greatly improves the performance of remove_atoms_from_image_using_2d_gaussian, making it easier to work with large images with multiple sublattices.
It also includes several minor style and linting fixes.
About Atomap
Atomap is a Python library for analysing atomic resolution scanning transmission electron microscopy images. It relies in fitting 2-D Gaussian functions to every atomic column in an image, and automatically find all major symmetry axes. The full procedure is explained in the article Atomap: a new software tool for the automated analysis of atomic resolution images using two-dimensional Gaussian fitting.
The easiest way to try Atomap is via Binder: introductory Jupyter Notebook. To install Atomap on your own computer, see the install instructions.
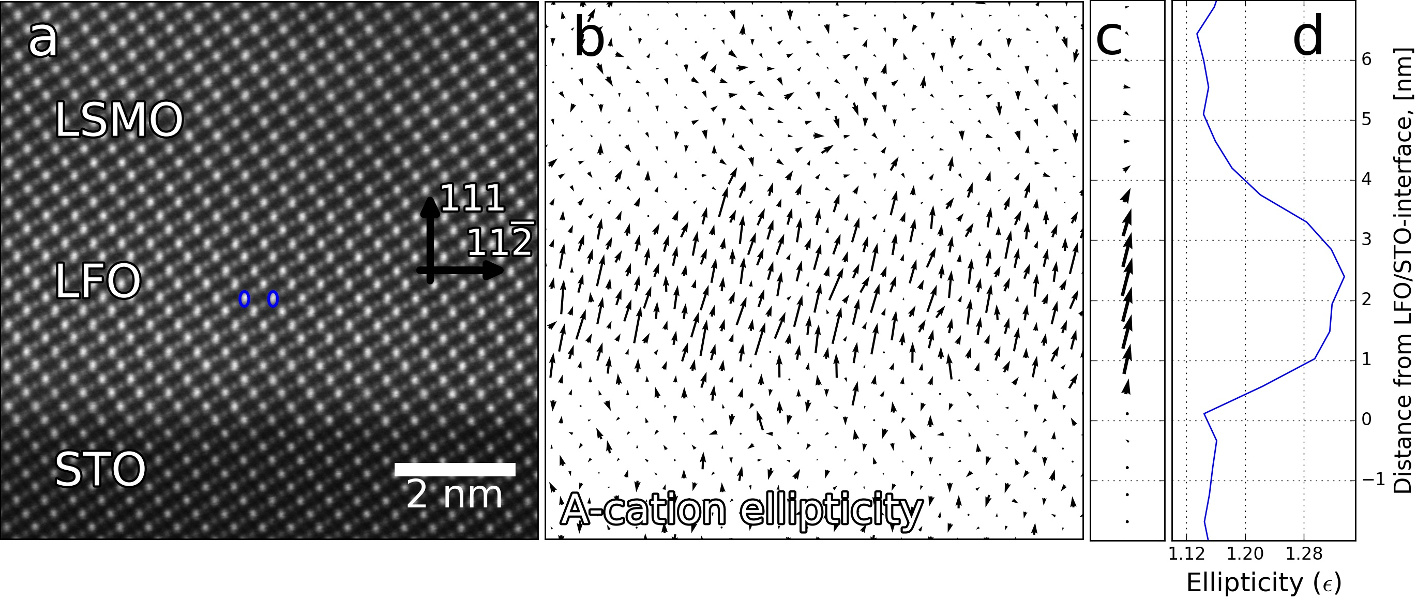
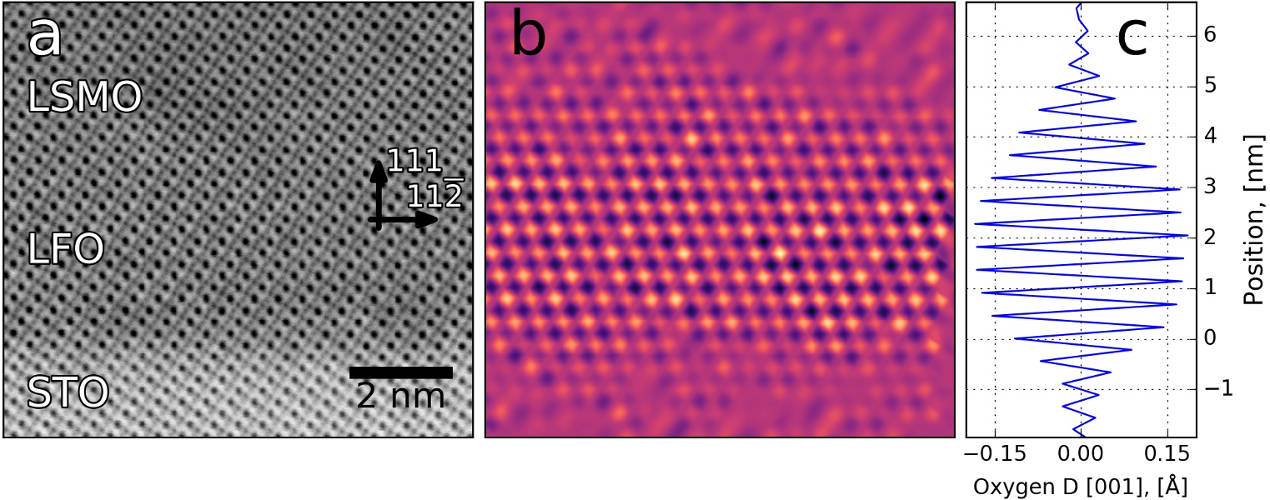
Mapping the variation in distance between oxygen columns. More information
Atomap is under development and still in alpha, so bugs and errors can be expected. Bug reports and feature requests are welcome on the issue tracker. Contributors are welcome too!
If you publish scientific articles using Atomap, please consider citing the article Atomap: a new software tool for the automated analysis of atomic resolution images using two-dimensional Gaussian fitting. (M. Nord et al, Advanced Structural and Chemical Imaging 2017 3:9)
Atomap is available under the GNU GPL v3 license. The source code is found in the GitHub repository.
Contents on this webpage
- Installing
- Start Atomap
- Finding the atom lattice
- Analysing atom lattices
- Dumbbell lattice
- Several phases
- GUI functionality
- Automation of the analysis
- Examples
- Quantification
- Working with atomic models
- Quantify scan distortions
- Making datasets for testing
- Plotting for publications
- Various tools
- Become a contributor
- Development guide
- API documentation
Old news
2024-10-20: Atomap 0.4.1 released
This is a maintenance release, making Atomap work with NumPy 2.0. It also updates Atomap to use pyproject.toml instead of setup.py.
2024-02-17: Atomap 0.4.0 released
This release is the first to work with the new 2.0.0 version of HyperSpy.
The major change is a large improvement in plotting speed, for example sublattice.plot() or sublattice.plot_planes(), thanks to a reimplementation of HyperSpy’s marker functionality. Thanks to Carter Francis for the merge request!
There has also been some minor updates of the install instructions, and fixes for deprecation warnings.
2023-11-18: Atomap 0.3.4 released
This release has a large performance improvement for the 2D Gaussian refinement, which is especially noticeable for large images with many atoms.
A new parameter, nearest_neighbors, has been added to sublattice.construct_zone_axes: increasing it will give more atomic planes.
Note: due to a new API-breaking version of HyperSpy (2.0.0), this Atomap release will be the last which will work with the older HyperSpy versions (1.x.x).
Thus, the next releases will only work with HyperSpy 2.0.0 or higher, most users should not notice any differences.
2023-5-17: Atomap 0.3.3 released
This is a small maintenance release, fixing some deprecation warning, style issues, and fixing a plotting bug when visualizing vectors.
2022-4-28: Atomap 0.3.2 released
This is a maintenance release, updating Atomap to work with the newly released HyperSpy 1.7.
2021-6-11: Atomap 0.3.1 released!
This is a small release, with one bug fix from Tom Slater, and two performance improvements from Niels Cautaerts. The latter are related to finding the initial atomic column positions, and will especially improve the processing time for large images.
2021-4-23: Try Atomap without having to install anything, via Binder
2021-1-30: Atomap 0.3.0 released!
Information about the 3D structure of
Sublatticescan now be added, allowing for 3D atomic models to be “converted” into anAtom_Latticevia the Atomic Simulation Environment, and vice versa. This 3D information can be set manually, or via the intensity for homogeneous structures see statistical method and working with atomic models for more info. Thanks to Tom Slater and Eoghan O’Connell for implementing this!The performance of atomic column integration has been greatly improved,
atomap.integrateandAtom_Lattice.integrate_column_intensitywill be much faster now. Thanks to Thomas Aarholt!set_scale()method inAtom_LatticeandSublatticehas been added for easily setting the scale and unit.Better support for visualizing
Atom_LatticeandSublatticeobjects with scaled units. Thanks to Eoghan O’Connell for adding this.
2020-1-29: Atomap 0.2.1 released!
New method for getting a pair distribution function, see the documentation for more info. Thanks to Tom Slater for implementing this!
Add a method for getting the local, often high frequency, scanning distortions utilizing the shape of the atomic columns: Quantify scan distortions.
Improved the progressbar when using Atomap in Jupyter Notebooks. Thanks to Alexander Skorikov!
2019-10-17: Atomap 0.2.0 released!
Greatly improved documentation, notebook and several new functions for analysing images with dumbbell features, like Si or GaAs.
New GUI tool for selection a subset of atom positions, making it easier to work on images with several phases.
Statistical quantification using Gaussian mixture model. Thanks to Tom Slater for implementing this!
Functions in the
Atom_Positionclass:calculate_max_intensity()andcalculate_min_intensity(). Thanks to Eoghan O’Connell for implementing this!
2019-03-05: Atomap 0.1.4 released!
This release includes:
Functions to find shifts within unit cells, often seen in materials with polarization. This can for example be used to find shifts in B-cations in relation to the A-cations in a perovskite structure: Finding polarization.
Big optimization in refining the atom positions using centre of mass, which makes it much easier to work with images containing a large number of atoms. For smaller images the improvement is around 5-10 times, while for larger ones it is 1000 times faster. Thanks to Thomas Aarholt for implementing this!
Similar optimizations to
get_model_image(), which is primarily used to generate test data via the Dummy data module. This means generating test data likeget_fantasite()is 60 times faster, with even bigger improvements to larger test data. Thanks to Annick De Backer at EMAT for tips on how to improve this!
Another optimization for the Integration of atomic columns functionality is in the pipeline, with a merge request from Thomas Aarholt.
2018-11-26: Atomap 0.1.3 released!
Major features in this release includes:
A GUI function for adding and removing atoms, which makes it easier to set up the initial atom positions. See Adding atoms using GUI for more information.
A GUI function for toggling if atom positions should be refined or not, see Toggle atom refine.
Better handling of hexagonal structures, by adding a adding a
vector_fractionparameter tofind_missing_atoms_from_zone_vector, thanks to Eoghan O’Connell for the suggestion!The addition of
mask_radiusto the refine functions, which makes it easier to work with non-perodic atom positions, for example Working with single atom sublattices.
2018-06-13: Atomap 0.1.2 released!
This is a minor release, including a signal attribute in Sublattice and Atom_Lattice classes, minor changes in package dependencies and some improvements to the documentation.
2018-03-25: Atomap 0.1.1 released!
The major new features are methods for integrating atomic column intensity and quantifying this intensity in ADF STEM images, see Integration of atomic columns and Absolute Integrator for more info. Thanks to Katherine E. MacArthur for adding this! Other features include tools for rotating atomic positions for plotting purposes (Rotating image and points), and reduced memory use when processing data with many atoms.
2017-11-16: Atomap 0.1.0 released!
We are happy to announce a new Atomap release. It includes a major makeover of the tutorial, start with Finding the atom lattice. New features in this release are methods for finding atomic column intensity, new and simple plotting tools and a module for generating test data.
2017-07-03: version 0.0.8 released!
New features: ability to process dumbbell structures, fitting of multiple 2D Gaussians at the same time, improved background subtraction during 2D Gaussian fitting, and processing of nanoparticles.